Sequence optimization
A module developer can classify a new module to 'Sequence optimization' when the module is relevant to sequence optimization such as codon optimization, UTR sequence optimization, or intron sequence optimization. To develop optimization algorithm, you will have to use a specialized class (class name: GeneConstruct) to access to a memory array of gene constructs and gene analysis table. In this section, we show a short demonstration how to develop 'Sequence optimization' module. For more details, check 'Class & module' section.
o Gene construct array and cell address
In the below figure, the first column ("#") represents gene constructs and each column includes information of a gene construct including gene component.
1. The first row means the first gene construct and each cell ("[Gene] CAI", "[Gene] Mismatched DNA", ..., or "[Gene] mRNA -> Structure") represents an individual analysis result of the first gene construct. Their cell addresses (row, column) are "(1,1)", "(1,2)", "(1,3)", ..., and "(1,7)", respectively. Please remember that gene construct number (#) is an essential index if you want to specify a certain gene construct to read and write sequence data.
2. For example, in case of "49.162", cell address is (4,3) where number 4 means 4th gene construct and number 3 means 3rd column.
3. In case of "40.1961", cell address is (6,5).
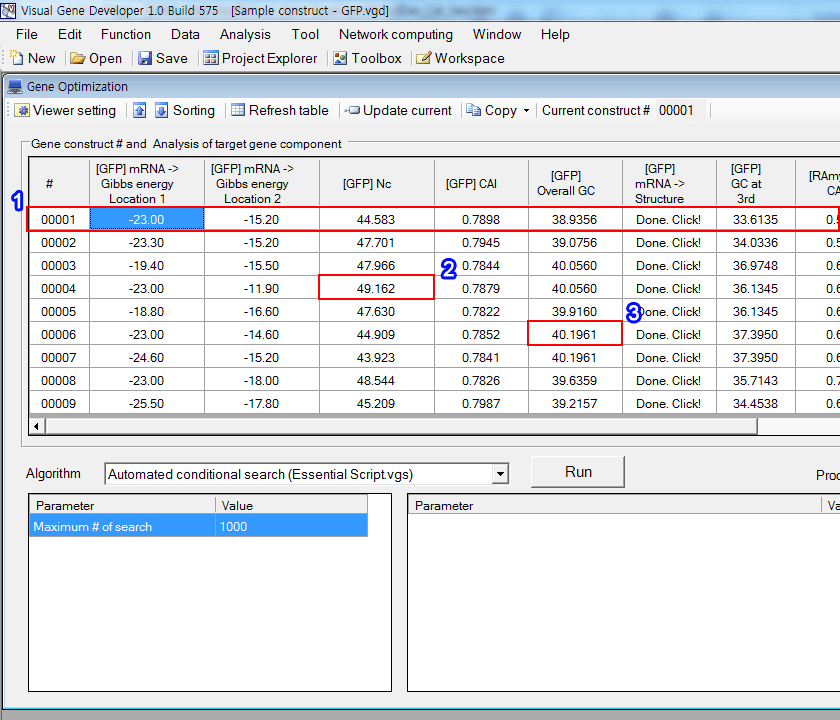
The cell address is also important when you access to the table.
o How to develop module
All modules are developed in the 'Module Editor' window
Step 1: Create module
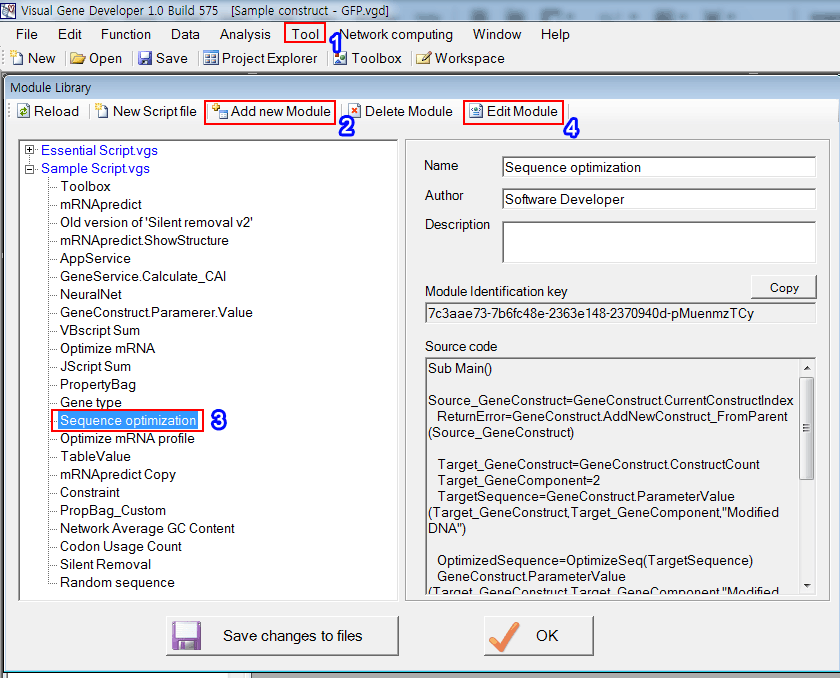
1. Click on the 'Module Library' in the 'Tool' menu
2. Click on the 'Add new Module' button to create a new module
3. Or, just choose the predefined module: 'Sequence optimization'
4. And then click on the 'Edit Module' button
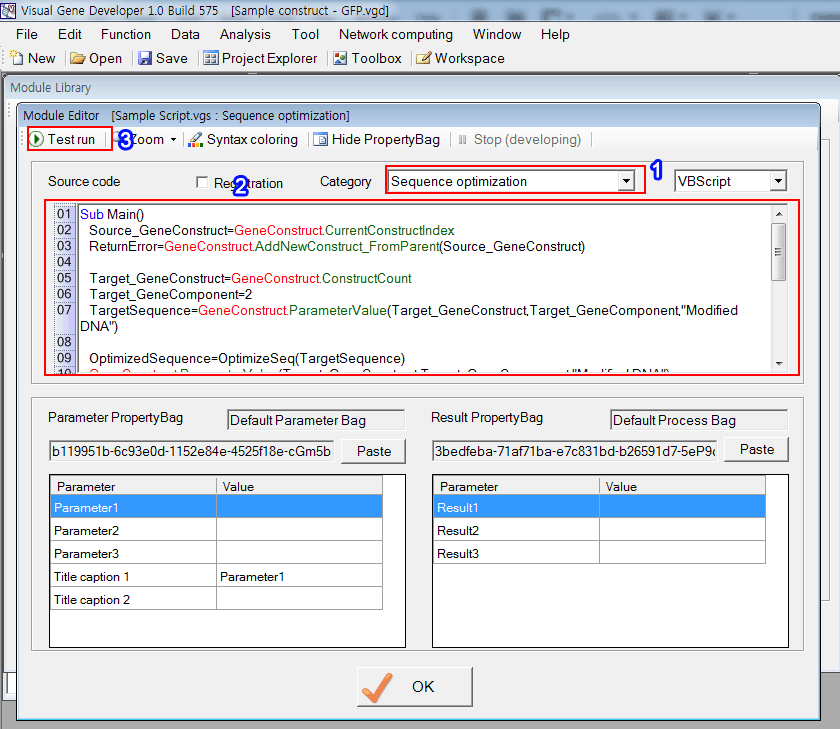
Step 2: Edit module
1. Choose 'Sequence optimization'
2. Check the example code
3. Click on the 'Test run' button to run the code
Here is a full source code (VBScript version)
|
Sub Main() ReturnError=GeneConstruct.Update_ConstructTable(Target_GeneConstruct)
'<----
Update table |
Overall function of the code is to copy a source gene construct to a new gene construct without modification.
Step 3: Test run
1. Choose 6th gene construct
2. Click on the 'Test run' button
3. A new gene construct will be shown in the 11th rows. In other words, the module copied the 6th gene construct to the 11th gene construct
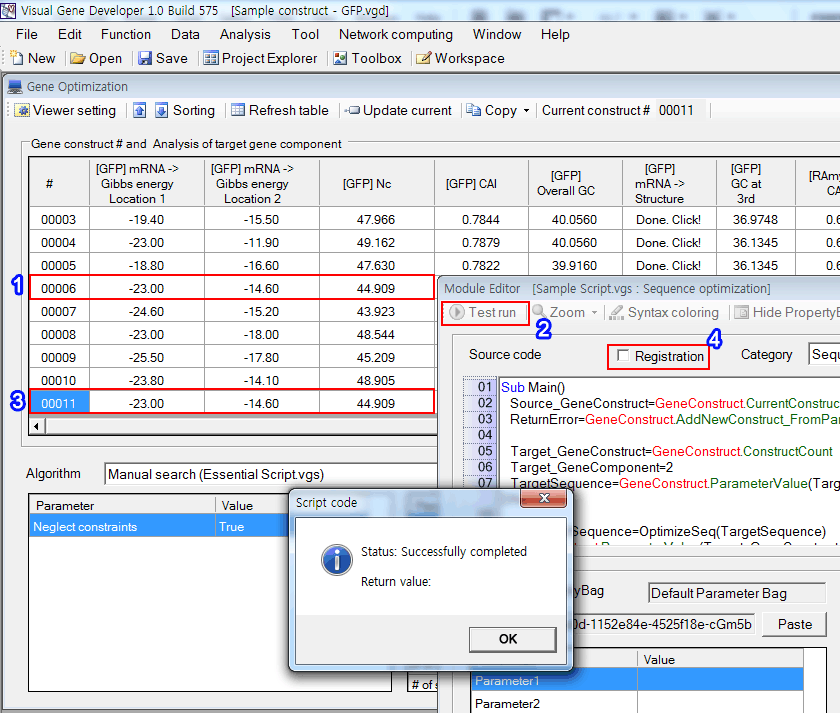
(4) To register the module to a proper location, check on 'Registration'.
Step 4: Use module
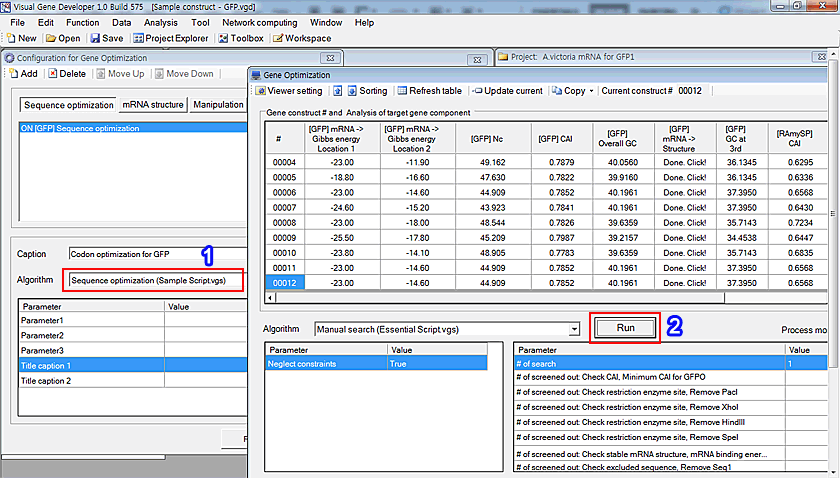
1.Once a module registered, you can see a new module in the 'Algorithm' selection box.
2. After adding 'Sequence optimization (Sample Script.vgs)', click on the 'Run' button to start optimization.
3. As shown in the Step 3, it will work the same way as 'Test run' button on 'Module Editor' window
o Specialize class: GeneConstruct
The class, GeneConstruct, contains many modules to access to the gene construct memory array and gene analysis table. For more details, check 'Class & module' section in the left menu.
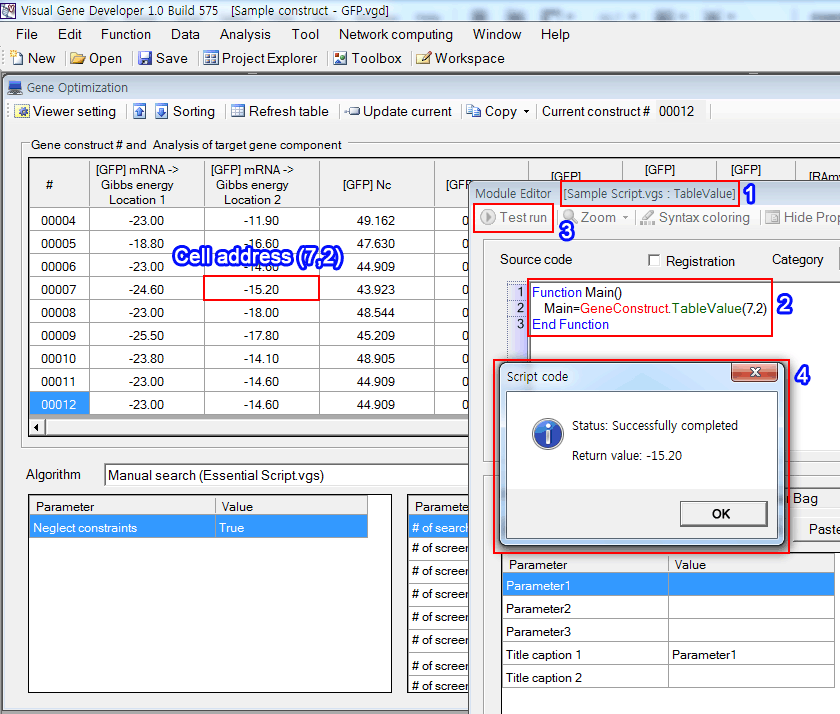
Here is an example that show how to read cell value of the Gene optimization table
1. Check the 'TableValue' module in the 'Sample Script.vgs' in 'Module Library' window
2. Review the source code. The 'GeneConstruct.TableValue(7,2)' means reading cell value at the address of (7,2) where 7 is the 7th gene construct.
3. Click on the 'Test run' button
4. The Message box shows the result.